In a recent study, researchers at the National Centre for Cell Science (NCCS), Pune have used Cryo-Electron microscopy to figure out the structure of an “orphan” receptor expressed in the central nervous system. The study provides important insights into the mechanisms via which this receptor functions, and sets a precedent for using Cryo-EM as a powerful tool for molecular investigation.

The field of structural biology received a significant boost when a new imaging technique emerged in the early 90s called Cryo-Electron Microscopy (Cryo-EM). It offered scientists a chance to photograph biomolecules and proteins in their natural molecular states. With the help of new ultrafast detectors used in such transmission electron microscopy, scientists can obtain unprecedented three-dimensional views of biological molecules. This opens the floodgates for not just explorative studies but also advanced targeted therapy.
Taking advantage of this cutting-edge tool, a team of Indian scientists, led by Janesh Kumar from the Laboratory of Membrane Protein Biology, National Centre for Cell Science (NCCS), Pune, provide the first insights into the structural architecture of a neurotransmitter-receptor protein called GluD1. The protein mediates critical nerve functions of the central nervous system like memory, mobility, cognition, and growth.
Nerve cells (neurons) are specialised cells that speak in an electrical language. They can propagate electrical signals at neuronal junctions called synapses with the help of specialised proteins called receptors. GluD1 is one such ionotropic (capable of forming ion channels) receptor residing in the cell membrane of neurons.
Usually, when information has to pass through a synapse, the transmitting cell membrane releases chemical messengers called ligands (or neurotransmitters) which act like ‘keys’ that fit into a pattern of ‘locks’ on a receptor protein on the receiving cell membrane. When ionotropic receptors like GluD1 bind the correct ligand, they open a pore in the cell membrane to facilitate the signal transfer.
However, GluD1 is a maverick: Although it belongs to a neurotransmitter-receptor family, it behaves like an ‘orphan’— neither binding nor getting activated by any known ligands to generate a passageway for the ions. GluD1’s modus operandi has baffled scientists for decades, further limited by the unavailability of an in-depth structural understanding of the molecule.
Now, Kumar and his team provide a ringside view of the elements that contribute to GluD1’s unique molecular attire and unravel its orphan-like behaviour. They observe that the molecular elements on GluD1 that actually perform the ‘linking’ at the site where the ligand binds to the ion pore are in a ‘relaxed’ state, which makes them ineffective in opening up a channel for ions in response to the ligand.
The Cryo-Electron microscope used for this study is an advanced, recently developed, high-power instrument used to photograph biomolecules. In this technique, the sample protein is flash-frozen (in its native form) at extremely low temperatures. Then, a beam of electrons is fired at the sample, which passes through the nano-sized specimen and gets scattered. A high-speed camera attached to the transmission electron microscope captures the scattered beam and takes rapid, high-resolution snapshots. The several thousand images that so emerge are sorted, integrated and computationally processed to obtain three-dimensional views of the protein molecules.
“Cryo-EM will impact and revolutionise structural biology in India,” says Vinothkumar Kutti Ragunath, Faculty- in- charge of the National Cryo-EM facility, Bangalore Life Science Cluster, NCBS, Bengaluru, who was not involved with this study. “In this technique, the molecules are imaged as single entities, and for difficult samples like that studied by Kumar’s team, it is extremely useful,” he adds.
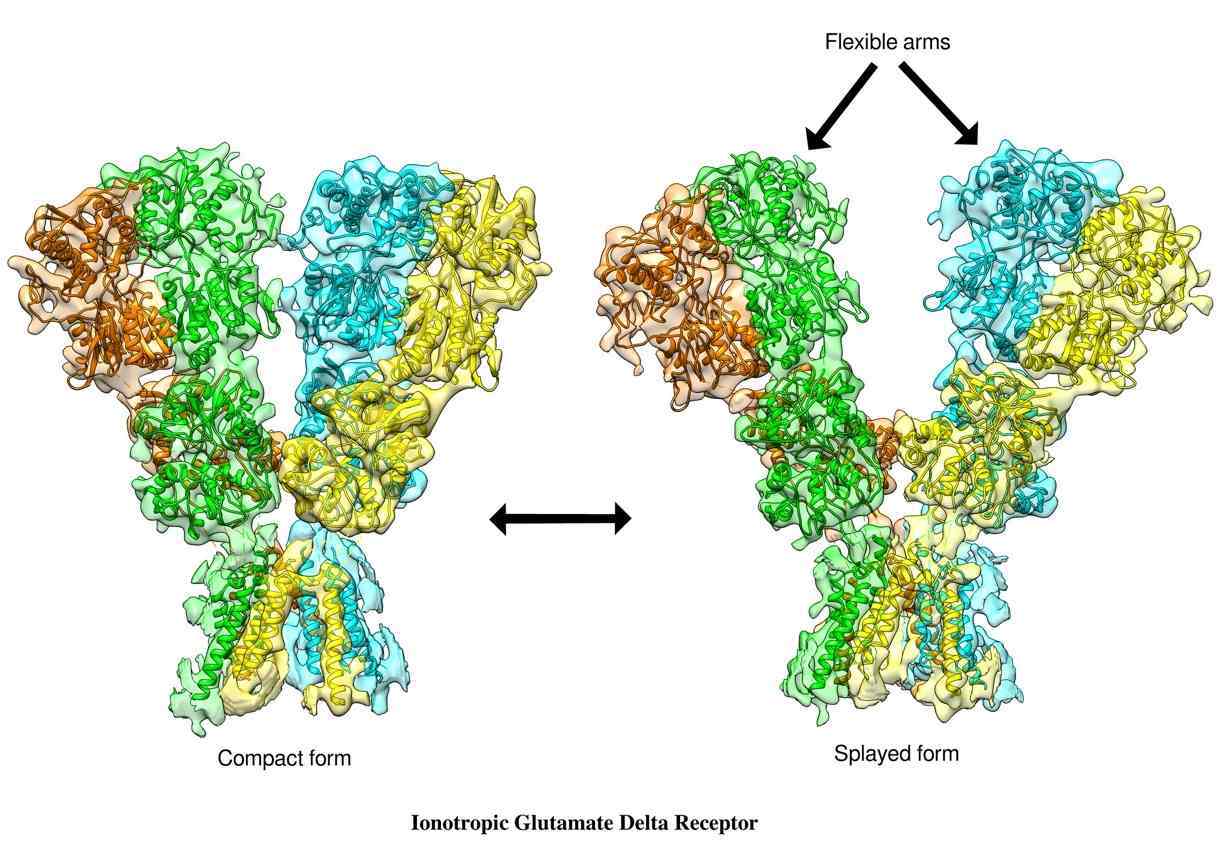
When the team deciphered the image of GluD1, they found it had a three-layered ‘Y’ shaped structure. The foot of the ‘Y’ remained buried in the cellular membrane, while the limbs of the ‘Y’ projected outside the cell. “The unique ‘Y’ shaped architecture adopted by GluD1 receptors was hitherto unknown in the glutamate receptor family. This unique shape gives clues into the ways GluD1 works,” says Kumar.
The molecular structure also gives a clear picture of precisely which part of the protein forms the ion pore, the shape and size of the protein chains that bundle together to form the gateway for ion transport, and the vestibule-like opening in the protein through which ions can pass through.
Kumar’s team extracted the protein from human cell cultures, a rigorous process. “Membrane proteins, in general, are challenging to work with as they express poorly in cells and are not stable once separated from the cellular membrane. GluD1 is prone to aggregation and precipitation even under slight changes in the extracellular environment,” explains Ananth Prasad Burada, first author of the study.
To overcome the hurdles, the researchers employed innovative chemical tweaks and alterations to existing protein extraction protocols. Following optimisation and slight protein modification, they found the right chemical conditions to stabilise GluD1 in laboratory conditions. “At each step, we conducted functional electrophysiological assays to ensure that the modifications don’t affect GluD1’s functional properties and the protein is physiologically relevant for imaging,” says Rajesh Vinnakota, a post-doctoral fellow at NCCS, and a co-author on the paper.
Functional proteins such as GluD1 are dynamic molecules, implying they change their shape to facilitate the ion flow. The study finds that calcium ions may play a role in the protein’s function; however, it is yet to be ascertained how effective this process is under physiological conditions.
Further investigations are underway to determine the exact triggers and factors that affect GluD1’s structure and function. “The detailed 3‑D maps provide a molecular blueprint of these important molecules which could be exploited for therapeutic targeting,” adds Kumar.